You are here : Home City of energies > HOME BIAM > Search > MEM > MEM research topics
Research topics
Bacterial radioresistance and oxidative stress sensing
By Laurence Blanchard and Arjan de Groot
Our research aims to decipher the molecular mechanisms that confer Deinococcus bacteria extreme resistance to radiation and DNA-damaging stresses (e.g. desiccation, UV, mitomycin C (antibiotic, anti-cancer agent), and oxidative stress in general). In this context, our work focuses on the characterisation of Deinococcus deserti, a radioresistant bacterium isolated from the Sahara desert, showing original characteristics – in particular compared to the model species Deinococcus radiodurans – highlighted by comparative proteogenomics (De Groot et al. (2009) PLoS Genet). The exploration of the biodiversity of Deinococcus, isolated from various environments around the world, is also at the heart of our current studies (Lim, Jung, Blanchard, de Groot (2019) FEMS Microbiol Rev).
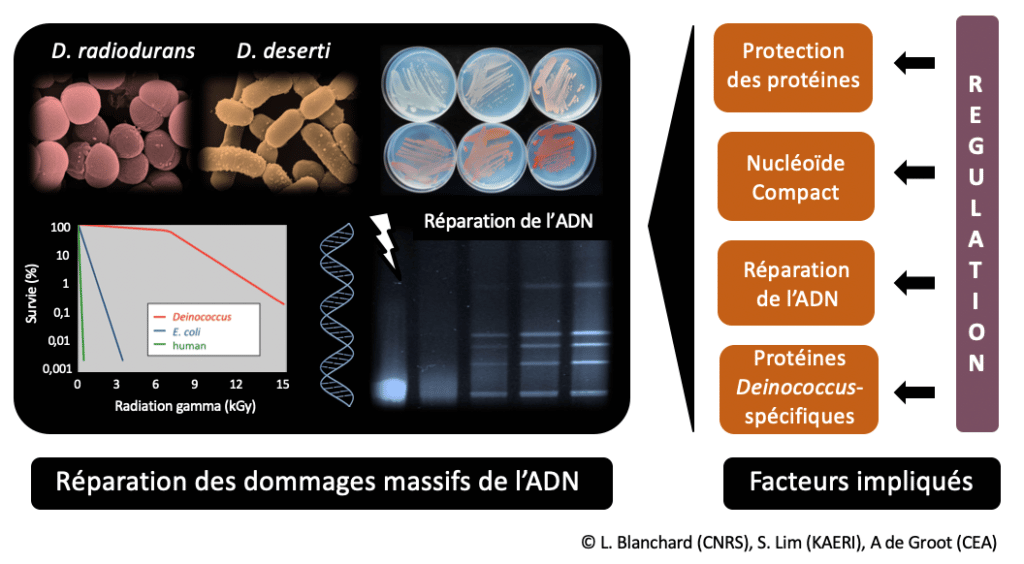
Deinococcus bacteria, cocci or rods, show coloured colonies on Petri dishes due to the presence of carotenoids. These bacteria tolerate much higher doses of ionising radiation than most bacteria and human cells (no loss of survival at 5,000 Gray (Gy) while a dose of 10 Gy is lethal to humans). This extreme resistance is related to the ability of Deinococcus to reconstitute an intact genome from a genome fragmented by irradiation, including efficient repair of many double-strand breaks that are lethal to most organisms (De Groot et al. (2009) PLoS Genet) (Figure 1).
Their secret, discovered step by step in recent years, lies in a combination of numerous finely regulated repair and protection factors and mechanisms (Lim, Jung, Blanchard, De Groot (2019) FEMS Microbiol Rev). Many mechanisms and functions of new proteins, however, remain to be deciphered.
Understanding how Deinococci resist radiation may lead to a better understanding of the radiation resistance of other organisms or cells (e.g. certain cancer cells with increased resistance to radiotherapy), or conversely to understanding why some cells are more radiosensitive than others.
Approaches used
The global, differential proteogenomic or transcriptomic (RNA sequencing) approaches we use allow the selection of genes/proteins related to radioresistance. The identified genes/proteins (e.g. radiation-induced, involved in DNA repair, specific to Deinococcus or D. deserti) are then characterized by genetic and structural biochemistry.
Main projects in progress
Within the ANR project NOVOREP, we are studying D. deserti (Cadarache) and D. radiodurans (coll. I2BC, Paris-Saclay). We focus on the SOS-independent stress response mechanism that we discovered, which is controlled by two key regulators, IrrE & DdrO (Figure 2) (Vujičić-Žagar et al., 2009 ; Ludanyi et al., 2014 ;Blanchard et al., 2017 ; De Groot et al., 2019 ; Magerand et al., 2021 ; De Groot & Blanchard, 2021). Following stress exposure, induction of several DNA repair genes occurs after cleavage of the XRE family transcriptional repressor DdrO by the separate COG2856 metalloprotease IrrE, which is essential for stress survival. We recently described that the peptidase function of IrrE is activated by increased zinc availability after stress, showing for the first time in prokaryotes a role for zinc as second messenger in redox signalling. We also solved the 3D structure of DdrO giving insight into the (de)repression mechanism.
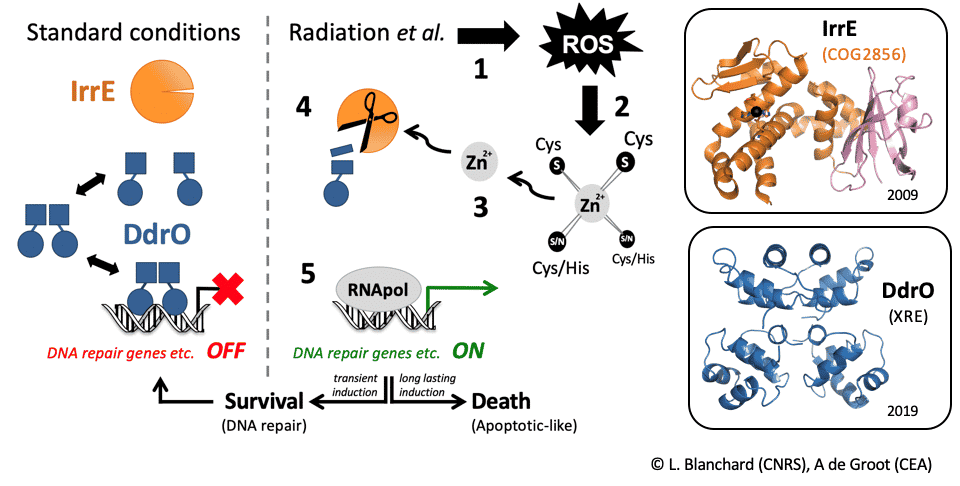
Figure 2: Description of a novel radiation response mechanism in Deinococcus. From signal to the fate of the cell: survival or programmed cell death
Proteins pairs similar to IrrE and DdrO are found in many environmental, pathogenic or industrially used bacteria, in which they control the production of phages or could control the production of toxins that induce dormancy and antibiotic tolerance in certain pathogens. The characterisation of these couples will provide an overview of the functioning of this new and large family of regulators, opening the way to the development of strategies to counteract the harmful effects of some of them in human health.
Exploring biodiversity is another axis. Our recent comparative analysis of 11 Deinococcus species revealed conservation and diversity of proteins involved in DNA repair and oxidative stress defence, and also highlighted various uncharacterized proteins (Lim et al., 2019). In the frame of a BIAM transversal project with Pascal Rey (PPV Team) and a Hubert Curien Program with our collaborators from KAERI South Korea, we have started to characterize a network of predicted thiol-reductases showing original features. This project also aims to create an international “Redox biology” consortium.
Metal Acquisition and Metalloproteins
By Pascal Arnoux
Essential metals (also called micronutrients) are vital resources for all living organisms. Metal homeostasis (which includes metal uptake, secretion and regulation of these functions) is therefore an important and fundamental process. Although there are general functional similarities, the underlying mechanisms are extremely diverse.
Perhaps the best example is the uptake of iron, which is relatively unavailable in many environments, especially in the presence of oxygen that oxidizes and precipitates iron. These conditions of non-bioavailability are also found at the interface between a host and a pathogen, a situation in which the host limits the availability of the metal precisely to prevent bacterial growth in a framework called “nutritional immunity.” Faced with this scarce and vital resource, bacteria adapt to synthesize siderophores with high affinity for iron, which they export to the extracellular space before the capture and import of the siderophore-Fe³+ complex. These siderophores are frequently associated with virulence and can also be used for the transport of metals other than iron, with few examples of other types of microbial metallophores.
In this context, in 2016 we described a new type of metallophore in pathogenic bacteria (Staphylococcus aureus and Pseudomonas aeruginosa ; see Figure below). These metallophores are related to nicotianamine, a plant metabolite used for long-distance uptake and transport of various micronutrients. In bacteria, these nicotianamine-like metallophores are primarily dedicated to zinc uptake in chelating environments, which are prevalent at the host-pathogen interface or in calcareous soils and some marine environments. Since these discoveries, we are pursuing this research by studying the diversity, regulation, role and evolution of nicotianamine-type metallophores in different bacterial models.
The Metallo team is organized around a PI (Pascal Arnoux), 2 technical and engineering supports (Catherine Brutesco and Géraldine Brandelet) and a PhD student (Manon Dassa-Valzer).
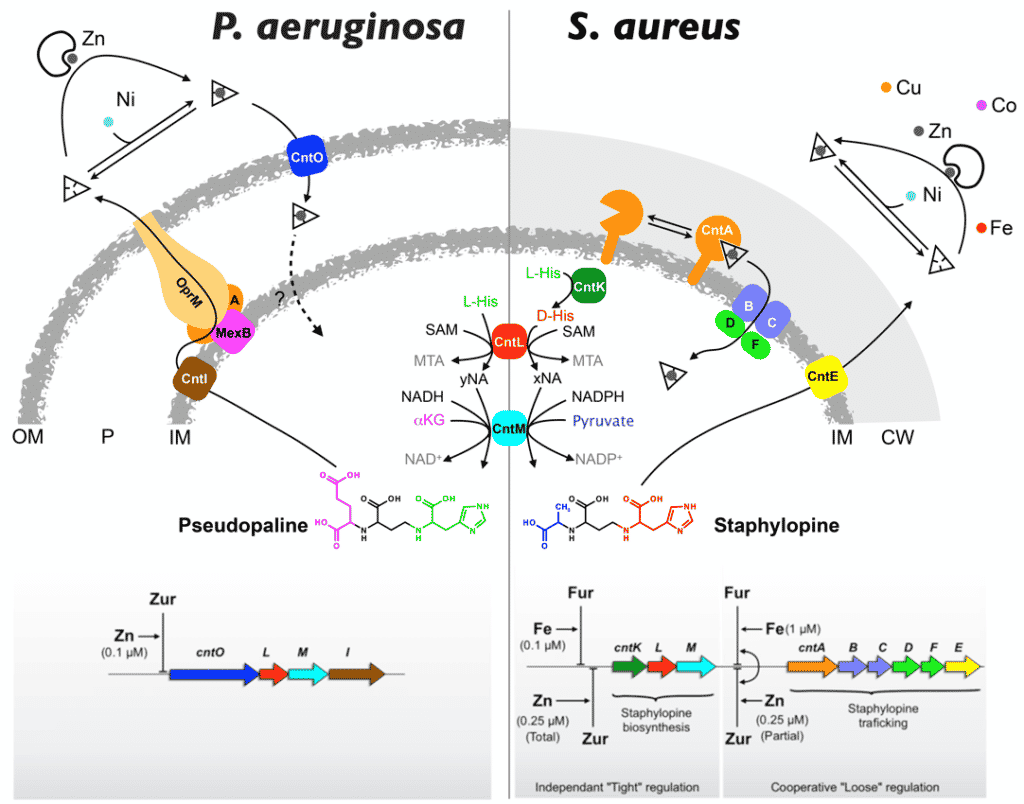
Figure 1 : Model of production and trafficking of nicotianamine-type metallophores in P. aeruginosa and S. aureus.
In addition to this main activity, I am also interested in the study of metalloproteins and their structural characterization. Current collaborations include the study of uranium binding to the UipA protein with V. Chapon, IPM, the characterization of an alkane-producing photoenzyme (APF) with D. Sorigue and F. Beisson, EBM in the BIAM Institute, and the characterization of a new family of molybdenum formate dehydrogenase with A. Walburger and A. Magalon, IMM, Marseille and FhtR, a heme sensor with D. Lechardeur, Micalis, INRAE, Jouy-en-Josas.
Microbial biomineralization (MicroBioMin)
MicroBioMin
Our group studies the mechanisms leading to the formation of biominerals by microorganisms and the ability of these microorganisms to navigate efficiently in complex environments
Our objectives encompass the understanding of the roles played by microorganisms in biogeochemical cycles and for the climate, the understanding of biological micro-swimmers and the development of effective micro-swimmers for various applications, as well as the respective advancement of bio- inspired and sustainable towards advanced materials. We are particularly interested in magnetotactic organisms forming ferrimagnetic crystals, and in coccolithophore algae forming calcium scales. Our approaches rely on the combination of in-house developed, mostly in situ, environmental and laboratory approaches. We use a combination of analytical techniques including microfluidics, electron and optical microscopy, X-ray spectromicroscopy, as well as omics
Our themes
1. Investigating biomineralization processes closer to native-state.
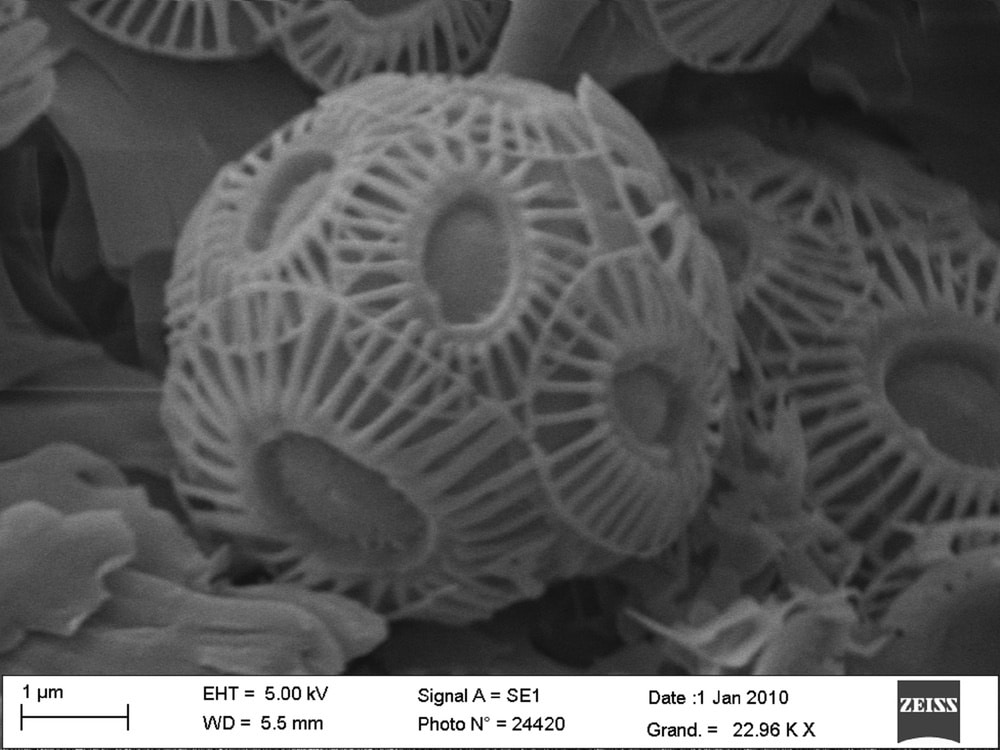

My research interests lie in the formation of inorganic nanomaterials and minerals via biomineralization processes, which involves biological molecules and templates produced by organisms that direct and control mineralization. Overall, revealing the necessary components and steps key to biomineralization is a means to adapt synthetic strategies to produce nano- and micro-structured materials with better control over morphology and size, and it is also a way to better understand the impact that biomineral formation has on our environment. Current projects seek to further our understanding of biomineralization processes by examining the formation of magnetite nanoparticles by magnetotactic bacteria (magnetosomes) and nanostructured calcite shells by coccolithophore microalgae (coccoliths) under native-state conditions (in liquid culture, in situ). This involves a combination of synchrotron-based X-ray spectroscopy and microscopy measurements, and microfluidic sample environments to capture biomineral formation within individual microorganisms. Studies are designed to provide unprecedented single cell-level analytics on the biomineralization process such as metal ion uptake, metal storage, biomineral formation rate, and identification of relevant precursors and intermediates.
2. (Bio)mineralization and Microswimmers: putting genes on the menu of materials science and soft condensed matter

Team manager
Key words
Genomics, bacterial genetics, molecular biology, microbiology, recombinant protein purification, biochemistry, structural biology, biophysics, enzymology.