You are here : Home City of energies > HOME BIAM > Search > PEPSS > PEPSS Research Themes
Research topics
Oxidative stress and non-stomatic responses to water stress
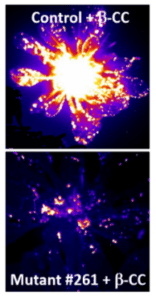

Many environmental stress conditions inhibit plant photosynthesis, leading to the production of reactive oxygen species in the chloroplasts and causing oxidation of macromolecules. We have identified oxidation products of the carotenoid β-carotene generated under photooxidative stress conditions, such as β-cyclocitral (β-CC) and β-cyclocitric acid (β-CCA), that act as signaling molecules triggering changes in nuclear gene expression and increasing plant stress tolerance (Havaux 2021). For example, watering Arabidopsis plants with a β-CCA solution increases their drought tolerance (D’Alessandro et al. 2019). This phenomenon is not attributable to a change in stomatal function, and our current work is focused on elucidating the β-CC(A)-induced signaling pathways and the mechanisms underlying the increased drought tolerance. In this context, we are trying to identify key genes in the β-CC(A) signaling pathways by combining transcriptomic analyses, genetic screens, and Arabidopsis mutant studies. We have already isolated a number of β-CC-unresponsive mutants and sequenced the genome of some of them, providing several candidate genes involved in β-CC signaling. In addition, a translational research work is in progress, in which the information collected in Arabidopsis is transferred to agronomic species, such as grapevine (in collaboration with two INRAE laboratories, SVQV Colmar and LPSE Montpellier). This work will provide the basis for the practical use of β-CCA as a phytoprotectant to improve drought tolerance in crops.
Stress-induced apocarotenoids in growth regulation and plant-plant communication
A major effect of β-CC(A) is to induce defense mechanisms at the expense of growth and development (Ramel et al. 2012). Accordingly, treatments of Arabidopsis plants with β-CCA were found to inhibit root growth. We identified genes that are induced by β-CCA and play a role in this root growth inhibition. The link to drought tolerance is currently being investigated through constitutive overexpression of these genes in Arabidopsis.
Unlike β-CCA, β-CC is a volatile compound that could potentially serve as a signal in plant-to-plant communication (Havaux 2021). This long-range signaling of β-CC and other volatile apocarotenoids is currently being studied in Arabidopsis and tomato under different environmental conditions including vegetation shade in intercropping agrosystems.
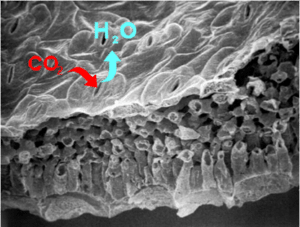
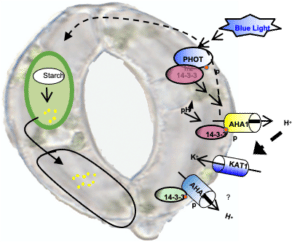
Using genetics, cell imaging, and electrophysiology approaches, we are exploring how ion fluxes are coordinated between membranes and cellular compartments in order to understand the mechanisms at the core of ion flux coordination during the opening and closing of stomata. These ion fluxes are crucial to the regulation of membrane potential and in signaling as well as in the turgor of cells. At the level of the plasma membrane of the guard cells, one of the major players in stomatal regulation is the activity of the H+ pumps, which create the electrochemical potential necessary for K+ to enter. In order to determine the roles of this family of transporters and their regulation in guard cells, several genetic resources are developed and exploited. Identifying and characterizing these fine mechanisms regulating the signaling pathways of guard cells is one of the original strategies for improving crop resistance to water stress.
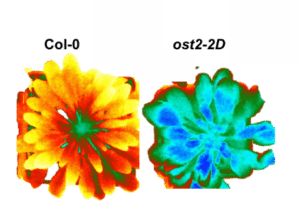
The role of H+-ATPases in Phytotechnology (PI: N. Leonhardt)

The pleiotropic roles of plasma membrane H+-ATPases are also illustrated by their impact on ion uptake via regulation of membrane potential or H+ efflux modulating the physicochemical properties of the rhizosphere. Based on the work performed under the PIA-DEMETERRES project (2016-2021), we have demonstrated that the mechanisms of cesium absorption by plants are affected by several parameters, including soil chemistry and plant physiology. The chemical properties of cesium are similar to those of potassium. It is generally accepted that Cs enters plants primarily through the K+ transport systems that provide nutrition to plants. The identification and characterization of these transporters are currently being conducted in collaboration with several academic and industrial institutions. Furthermore, the rhizospheric parameters at the soil/root interface as well as their modification by the root system play a crucial role in the absorption of Cs. In particular, the architecture, plasticity and transport activity of roots can modify the soil pH affecting the bioavailability of cesium and its uptake by plants. In this context, the identification and modification of the activity of the main proton pumping ATPases are currently being studied in order to increase (phytoremediation) or limit (phytostabilization) the absorption of cesium by modulating the availability of Cs in the rhizosphere and/or by regulating root transport systems for Cs uptake.

https://www.youtube.com/watch?v=LzO-I5DfL5U
Team manager: Nathalie PRAT
Key words
Arabidopsis thaliana; Cell biology; Genetics; Metals; Stomata; Root Architecture racinaire; Transporters; Signal transduction; Phosphate; Plant Physiology; QTL; Growth; Root, Water stress


